Consider a fragment of a protein chain from amino acid a to b, for example, a loop. Modeling fluctuations of the fragment under physiological conditions requires finding conformations where the fragment termini amino acids remain connected to the rest of the protein. This problem, often referred to as loop modeling or loop closure for fragment loops, poses the need to compute geometrically-constrained conformations of a fragment of the protein chain that result in low-energy conformations of the entire protein chain. This is illustrated below for the 12-aa loop of CI2.
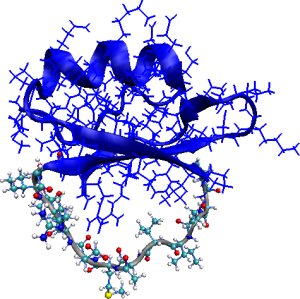
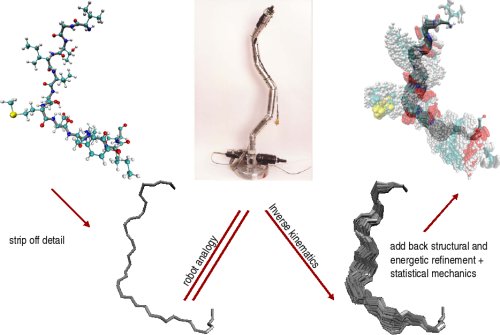
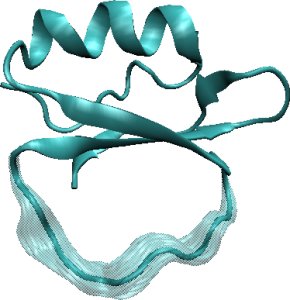
Left: Loop fragment of 12 amino acids (in grey) is constrained to connect to the rest of the CI2 protein structure (in blue). Center: The main steps of FEM. Right: The FEM method allows computing low-energy geometrically-constrained conformations of the loop fragment in CI2.
The Fragment Ensemble Method (FEM) was developed to obtain an ensemble of physical conformations for a protein fragment. FEM first strips a fragment off its side chains to model the fragment backbone as an open kinematic chain. This analogy is exploited to sample backbone conformations similarly to sampling configurations of a kinematic chain. An optimization-based inverse kinematics method (Cyclic Coordinate Descent) is applied to each sampled conformation in order to obtain closure conformation; that is, fragment conformations that satisfy the termini constraints.
Low-energy side-chain configurations are then placed on each backbone conformation. Energetic refinement is conducted to reduce unfavorable inter-atomic interactions. The refinement focuses mostly on the fragment while allowing small fluctuations in the rest of the protein structure. Following a statistical mechanics framework, each resulting conformation is weighted by its Boltzmann probability, which provides a quantitative measure of feasibility under physiological conditions.
The FEM method has been applied to characterize the flexibility of mobile protein fragments such as loops. In addition to its ability to reproduce the preference of strongly stable loop fragments for one average native conformation, the method can also model missing loops. The method has been applied to rigid and flexible loops of diverse lengths, from 12 to 31 amino acids, that reside on the surface or interior of protein structures.
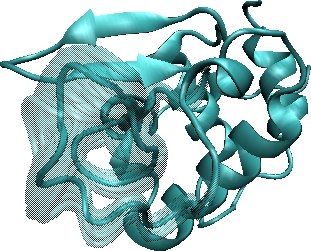
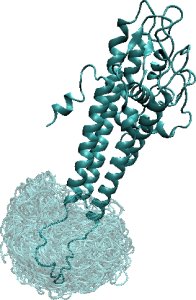
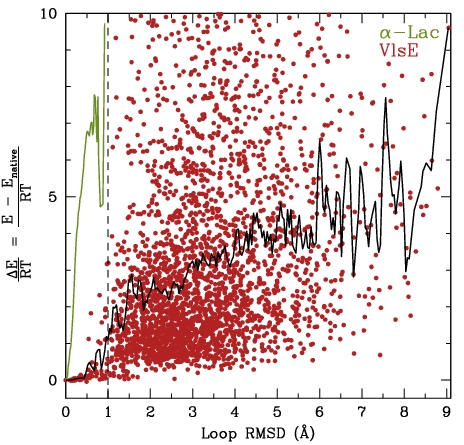
Left: The two ensembles show FEM-generated low-energy conformations for the 31-aa loop in alpha-Lac and 20-aa missing loop in VlsE. The computed conformations are shown in transparent, whereas the lowest-energy conformation is drawn in opaque. Right: The energy landscapes of FEM-generated ensembles for the alpha-Lac and VlsE loops show that the loop in alpha-Lac prefers one conformational state, while the missing VlsE loop is practically disordered.
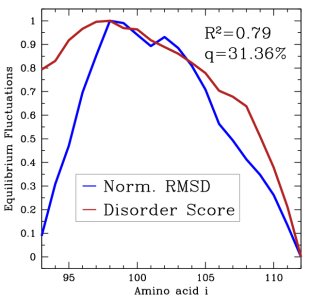
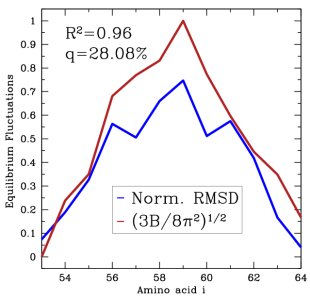
Left: Disorder scores measured as Boltzmann averages over generated VlsE loop ensemble agree well with scores predicted from the loop sequence. Right: B factors measured as Boltzmann averages over generated ensemble for CI2 loop agree well with B factors available from X-ray experiments.
This work appears in: 1) Amarda Shehu, Cecilia Clementi, and Lydia E. Kavraki "Sampling Conformation Space to Model Equilibrium Fluctuations in Proteins" Algorithmica, 2007, 48(4):303-327; and 2) Amarda Shehu, Cecilia Clementi, and Lydia E. Kavraki "Modeling Protein Conformational Ensembles: From Missing Loops to Equilibrium Fluctuations" Proteins: Structure, Function, and Bioinformatics 2006, 65(1):164-179.
On this Project:
-
Amarda Shehu
Cecilia Clementi
Lydia Kavraki
This project is completed.

